-Search query
-Search result
Showing 1 - 50 of 8,342 items for (author: shi & y)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
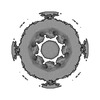
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-60561: 
Cryo-EM structure of uropathogenic Escherichia coli CysK:CdiA:tRNA complex A
Method: single particle / : Feng Z, Yashiro Y, Tomita K

EMDB-60562: 
Cryo-EM structure of uropathogenic Escherichia coli CysK:CdiA:tRNA complex B
Method: single particle / : Feng Z, Yashiro Y, Tomita K

EMDB-60563: 
Cryo-EM structure of uropathogenic Escherichia coli 2:1 CysK:CdiA complex
Method: single particle / : Feng Z, Yashiro Y, Tomita K

EMDB-60564: 
Cryo-EM structure of uropathogenic Escherichia coli 2:2 CysK:CdiA complex
Method: single particle / : Feng Z, Yashiro Y, Tomita K
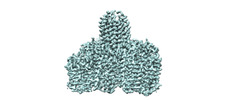

EMDB-18162: 
N5-methyl-H4MPT:CoM methyltransferase -coenzyme M complex + CoM
Method: single particle / : Aziz I, Vonck J, Ermler U

EMDB-42487: 
Cryo-EM reconstruction of Staphylococcus aureus oleate hydratase (OhyA) dimer of dimers
Method: single particle / : Oldham ML, Qayyum MZ

EMDB-37571: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37572: 
cryo-EM structure of alligator haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37573: 
cryo-EM structure of alligator haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37574: 
cryo-EM structure of human haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37575: 
cryo-EM structure of human haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37576: 
cryo-EM structure of human haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wix: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wiy: 
cryo-EM structure of alligator haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wiz: 
cryo-EM structure of alligator haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wj0: 
cryo-EM structure of human haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wj1: 
cryo-EM structure of human haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wj2: 
cryo-EM structure of human haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-45005: 
Paired Helical Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D

EMDB-45007: 
Straight Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D

EMDB-45008: 
Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D

EMDB-45009: 
Straight Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D
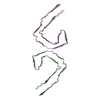
PDB-9bxi: 
Paired Helical Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D
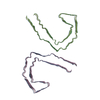
PDB-9bxo: 
Straight Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D
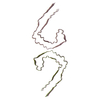
PDB-9bxq: 
Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
